Sat Aug 30 14:15:13 PDT 2008
Need to Know the Structure of a Molecule?
If you ever need to know the structure of a given molecule, use the eMolecules (http://www.emolecules.com) site to obtain all the information that you need.
I have found this site extremely useful for obtaining chemical information rapidly. The site has a large database of chemical information, obtained from the suppliers of a range of chemicals and chemical intermediates. You can query the database using an effective Java sketching tool, by SMILES string, or by common names.
Once you have obtained your hit you will see the various suppliers of the compound, so if you are a chemist you will be able to buy the appropriate amount, you will also be able to obtain additional information on the compound. The additional information includes the SMILES string, and using the SMILES string you can obtain a chemical connectivity file (.mol file) from the InChi site: http://inchi.info/converter_en.html.
So, if you are interested in the structure of Aspirin, for example, head over to eMolecules to obtain a SMILES string for Aspirin (e.g., CC(=O)Oc1ccccc1C(=O)O), then go to the InChi site, to obtain a .mol file. Then you can read the .mol file into PyMol, or your favorite molecular viewer, to see the structure of Aspirin.
These sites will give you a good idea of the structure of a range of molecules. The procedure above does not include a treatment of the lower energy confirmation of the shape of the molecule, and it does not include adding hydrogen atoms to the molecule. I will cover those steps in a future posting.
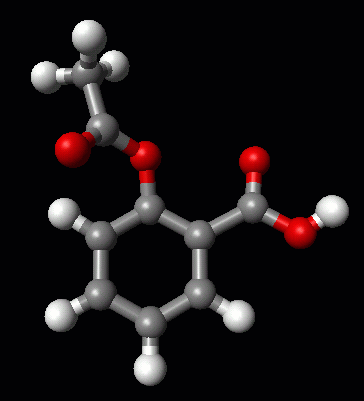
The Structure of Aspirin