| Su | Mo | Tu | We | Th | Fr | Sa |
|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 |
| 8 | 9 | 10 | 11 | 12 | 13 | 14 |
| 15 | 16 | 17 | 18 | 19 | 20 | 21 |
| 22 | 23 | 24 | 25 | 26 | 27 | 28 |
| 29 | 30 | 31 |
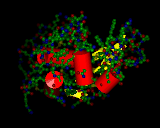
The molecule above is oxycodone, the molecule that very likely killed Prince.
The shape and chemistry of oxycodone are very similar to morphine and heroin.
Although oxycodone blocks pain and produces euphoria it is very addictive. It is hard to stop taking oxycodone because of the painful withdrawal symptoms that lack of the drug produces.
Like morphine and heroin, oxycodone binds to receptors which are normally the home of peptide hormones. The peptide hormones are the neurotransmitters which make humans feel good, scared, hungry, or satisfied. Such receptors have a hard time dealing with generally rigid molecules like oxycodone. Such molecules are not broken down like flexible peptide neurotransmitters and exert their influence on the receptor for extended periods of time.

|
The image on the left shows the details of a galaxy simulation of the style described by Schulman and Seiden.
The galaxy is composed of rings divided into cells with equal areas. Initially a random selection of galaxy cells are made active. Active galaxy cells are shown as being occupied by a black circle. After a galaxy cell has been active for a time step it is no longer considered active but a steadily diminishing circle is drawn in that cell to show that it was recently active.
With each simulation step, the cells in the vicinity of active cells are examined. These neighboring cells are made active themselves with a probability of 0.18. As there are 6 nearest neighbor cells for each active cell, this gives a slight bias in favor of more active galaxy regions with each timestep (6*0.18=1.08), which is counterbalanced by the fact that sometimes a given cell may be triggered by two neighboring active cells. Each cell is only active for one time step.
The rings of the galaxy rotate around the galaxy center. The speed of each rings rotation is set so that the rings rotate withe a constant velocity, not a constant angular velocity. So the outer rings perform fewer rotations per unit time than the inner rings.
The basic model is then one that assumes that the interstellar medium rotates at a constant velocity and that star formation is triggered by supernovae. Each supernova explosion can set off star formation in adjacent regions of the galaxy with a probability of 0.18. This is because the entire galaxy is composed of a dilute collection of hydrogen atoms. Given a suitable trigger, such as a nearby supernova, enough hydrogen can be compressed by the shockwave to trigger the gravitational build up of larger amounts of hydrogen leading to the formation of a star.
This simple model, constant rotational velocity, and adjacent star formation with a defined probability, leads to spiral galaxy formation. That might be a coincidence or it might indicate that the basic physics of galaxy formation is captured by this nearest neighbor model!
For additional information on galaxy simulation, see the following links on the galaxy simulation computer program and a video of galaxy formation.
I decide to put this site's directories under revision control so that I can maintain the history of changes with ease. Here are the commands that I used.
First I created a simple subversion repository based on the file system. This one is called 'svn' - which is not too sophisticated...
svnadmin create --fs-type fsfs svn
Then create a trunk directory in this new repository.
svn mkdir file://`pwd`/svn/trunk -m 'Creating trunk'
Then I imported the bin and public_html directories of the site into the new repository.
svn import bin file://`pwd`/svn/trunk/bin -m 'Initial import of bin directory' svn import public_html file://`pwd`/svn/trunk/public_html -m 'Initial import of public_html directory'
Now the bin and public_html directories are in subversion the originals can be moved else where and the svn controlled versions checkout.
mkdir tmp mv bin tmp mv public_html tmp svn checkout file://`pwd`/svn/trunk/bin svn checkout file://`pwd`/svn/trunk/public_html
Now I can work away in the bin and public_html directories and when appropriate commit the changes to subversion with suitable comments so that they can be backed out or learnt from if necessary.
|
As shown on the left, I spent a little time today working out how to make the images on this site expandable. Here is what I came up with.
To do this, I put an onlick javascript handle on to the img element for a particular image, likel this:
<img src="lysozyme_scale.gif" alt="(Lysozyme)" onclick="toggle(this)" />
The toggle function uses the supplied image file name to determine if it should display the 'full' image or the scaled down image (which I call lysozyme_scale.gif in this case). Here is the javascript:
function toggle(image) {
if (image.src.match("scale")) {
image.src = image.src.replace(/_scale/,"");
} else {
var ext = image.src.slice(-4);
image.src = image.src.slice(0,-4);
image.src = image.src.concat("_scale");
image.src = image.src.concat(ext);
}
}
...and the javascript file which contains this function has been added to the head section of every page which needs to display images.
<script type="text/javascript" src="./styles/scripts.js"></script>
To insure that each image provides information to the user about clickability - I put each image element into <a href="#"> </a> pair.
Overall this gives clickable images which expand to provide the user with additional visual information. This is convenient and the necessary modifications are small. One simply needs to follow the naming convention of filename_scale.png for the smaller image. So this seems to be both useful for users and convenient for the website maintainer.
Here is the version of the galaxy program used to generate the movies of the last few days.
#include <stdio.h>
#include <stdlib.h>
#include <math.h>
// PROGRAM galaxy
// model proposed by Schulman and Seiden
//
// ZFS - changed so that graphical output is handled in postscript
// included a minor fix to avoid stomping on memory and setting the time
// to 15, and increased the number of comments
//
// Usage
//
// Compile with:
// gcc galaxy.c
//
// Run with:
// ./a.out
//
// You will generate a collection of postscript files which show the
// evolution of your model galaxy
int main();
void parameter();
void initial();
void evolve();
void create(int r, int a, int *newactive, int newactive_r[], int newactive_a[]);
void plot_spiral();
float rand1();
// if printout is set to a positive integer this value is used to name
// the output postscript file
int printout=0;
int active_a[10001];
int active_r[10001];
int cell[51][301];
int dt;
int nactive;
int nring;
double p;
double s;
int t;
double two_pi;
double v;
double pi;
int main()
{
long int i;
parameter();
initial();
for(i=1;i<=1000;i++){
evolve();
printout=i;
plot_spiral();
t += dt;
}
return 0;
}
void parameter()
{
pi=3.141593;
nring = 50; // number of rings
nactive = 200; // number of initial active cells
v = 1; // circular velocity
p = 0.18; // star formation probability
dt = 10; // time step
s = 2*pi/6; // cell width
two_pi = 2*pi;
}
void initial()
{
int a, i, r, x, y, Itmp1_, Itmp2_;
float xwin, ywin;
float xtmp, ytmp;
for(Itmp1_ = 0; Itmp1_ <= 50; ++Itmp1_)
for(Itmp2_ = 0; Itmp2_ <= 300; ++Itmp2_)
cell[Itmp1_][Itmp2_] = 0;
// randomly activate cells
// this is done by choosing a ring by choosing points evenly on a grid
// and determining which ring the points lie in (to avoid overly sampling
// the small rings. The angle (a) about the ring can be chosen evenly for
// each ring.
i = 0;
do
{
do
{
x = (int)(nring*rand1()) + 1;
y = (int)(nring*rand1()) + 1;
xtmp=(float)x;
ytmp=(float)y;
r = (int)sqrt(xtmp*xtmp+ ytmp*ytmp) + 1;
} while(!(r <= nring));
a = (int)(6*r*rand1()) + 1; // array index corresponds to an angle
if(cell[r][a] == 0)
{
++i;
active_a[i] = a; // location of active region
active_r[i] = r;
// activate region, stars live for 15 time steps
cell[r][a] = 15;
}
} while(!(i == nactive));
t = 0; // initial time
}
void evolve()
{
int a, am, ap, i, newactive, newactive_a[10001], newactive_r[10001], r,
Itmp1_;
double angle, wt;
// number of active star clusters for next time step
newactive = 0;
for(i = 1; i <= nactive; ++i)
{
r = active_r[i];
a = active_a[i];
create(r, a+1, &newactive, newactive_r, newactive_a);
create(r, a-1, &newactive, newactive_r, newactive_a);
// angle is the angle of the current region, the angles of the next larger
// ring adjacent clusters are ap and next smaller ring clusters are am
//
// these angles are computed based on the ring velocity (which depends on the
// ring in question), the simulation time, and the overall velocity setting
angle = fmod((double)(a*s + v*t)/r, two_pi);
if(r < nring) // activate cells in next larger ring
{
wt = fmod((double)v*t/(r+1), two_pi);
ap = (int)((angle - wt)*(r+1)/s);
ap = ((ap) % (6*(r+1)));
create(r+1, ap, &newactive, newactive_r, newactive_a);
create(r+1, ap+1, &newactive, newactive_r, newactive_a);
}
if(r > 1) // activate cells in next smaller ring
{
wt = fmod((double)v*t/(r-1), two_pi);
am = (int)((angle - wt)*(r-1)/s);
am = ((am) % (6*(r-1)));
create(r-1, am, &newactive, newactive_r, newactive_a);
create(r-1, am+1, &newactive, newactive_r, newactive_a);
}
}
// copy the new actives into the actives lists - ready for the
// next time step
nactive = newactive;
for(Itmp1_ = 0; Itmp1_ <= 10000; ++Itmp1_) {
active_r[Itmp1_] = newactive_r[Itmp1_];
active_a[Itmp1_] = newactive_a[Itmp1_];
}
}
void create(int r, int a, int *newactive, int newactive_r[], int newactive_a[])
// create star clusters
{
if(a < 1)
a += 6*r;
// The original program had a bug in that a could be 301 on occasion (for 50th
// ring) and this then set the simulation time to '15' by reaching beyond the
// end of the cell matrix...this problem is avoided by the following test which
// imposes a circular boundary on the angle in the positive a direction (as
// was originally done in the negative a direction
if(a > 300)
a -= 6*r;
// Now see if a supplied region should become active because of its
// currenly proximal active region - this happens with a probability 'p'
if(rand1() < p && cell[r][a] != 15)
{
++(*newactive);
newactive_a[(*newactive)] = a;
newactive_r[(*newactive)] = r;
cell[r][a] = 15; // activate cell
}
}
void plot_spiral()
{
int a, r;
double plotsize, theta, x, y;
double theta2;
double scale=4.0;
double xoff=320.0;
double yoff=475.0;
char filename[20];
FILE * fd;
// Here we output postscript disks with radii which correspond to the
// age of a given star cluster. The location of a given cluster is
// determined based on the current simulation time and the velocity of a
// given ring.
sprintf(filename, "%07d.ps",printout);
if(printout){
fd=fopen(filename, "w");
fprintf(fd,"%%!PS\n");
fprintf(fd,"%%%%BoundingBox: 0 0 640 892\n");
fprintf(fd,"newpath\n");
fprintf(fd,"0.1 setlinewidth\n");
fprintf(fd,"0.0 0.0 0.0 setrgbcolor\n");
}
for(r = 1; r <= nring; ++r)
{
for(a = 1; a <= 6*r; ++a)
{
if(cell[r][a] > 0)
{
theta = (double)(a*s + (double)v*(double)t)/(double)r;
theta2 = fmod(theta, two_pi);
theta = theta2;
x = r*cos(theta);
y = r*sin(theta);
plotsize = cell[r][a]/15.0;
if(printout){
fprintf(fd, "newpath\n");
fprintf(fd, "%f %f\n", x*scale+xoff,y*scale+yoff);
fprintf(fd, "%f 0 360 arc closepath\n", 1.0*plotsize*scale);
fprintf(fd, "0.0 setgray fill\n");
fprintf(fd, "stroke\n");
}
// reduce star clusters lifetime
--(cell[r][a]);
}
}
}
if(printout){
fprintf(fd,"showpage\n");
fclose(fd);
}
}
// the code assumed that rand() returned a real number between 0 and 1,
// hence the following function which does this.
float rand1()
{
return((float)rand()/(float)RAND_MAX);
}
I found the original source code from here ... which comes from this compendium of code here from the book by Harvey Gould and Jan Tobochnik, 'Introduction to Computer Simulation Methods'. I have the first edition of this book (it has two volumes) and it is a really fantastic resource for those of us that did not pay enough attention in school but are interested in learning how physics operates.
I found a small problem with the original galaxy simulation. The program had the ability to set the simulation time to '15' under certain circumstances. The effect of that problem was to randomize the star clusters. I will post a link to the updated program in due course. The corrected animation is pasted above.
The movie above is a simulation of the formation of a spiral galaxy, based on a program described by Gould, Tobochnik, and Christian in their book 'An Introduction to Computer Simulation Methods'. Each of the cells in the simulation (which roughly correspond to the individual blobs that you can see in the movie) is a cluster of stars and each cluster is about 300 light years across. So this is a large simulation! This entire galaxy is about 30,000 light years in diameter which is not particularly large for a spiral galaxy, but it is quite a large system to simulate in a few seconds on a laptop - the complete simulation took about 100 seconds. (Making the movie took a bit longer...)
The basis for the calculation is the fact that an exploding star has a good chance of setting up the conditions for star formation in its vicinity, through the explosive shockwave that it sends out into space. This is coupled with the notion that the constituent rings of the galaxy move at constant velocity. With these assumptions the sprial and globular form of the galaxy emerges and evolves with time. (Like the distances, the time scales are vast, each timestep in the simulation is 107 years.)
You can perform very much more detailed simulations of galaxy formation, should the fancy strike you. But even this relatively simple model allows you to test the key facts and forces which are sufficient to explain the shape of spiral galaxies. Armed with the model, and its parameters, you can set about clarifying the key features which determine the different shapes of real galaxies, and even, if you are ambitious, the chances of the galaxy calming down and ceasing to produce new stars.
It turns out that the long range structure of the galaxy can be explained with only nearest neighbor interactions (if by nearest neighbor you mean cells about 300 light years across).